Something different?
Read about our new look website

Want to find out what documents we publish and how to access them?
Find out more
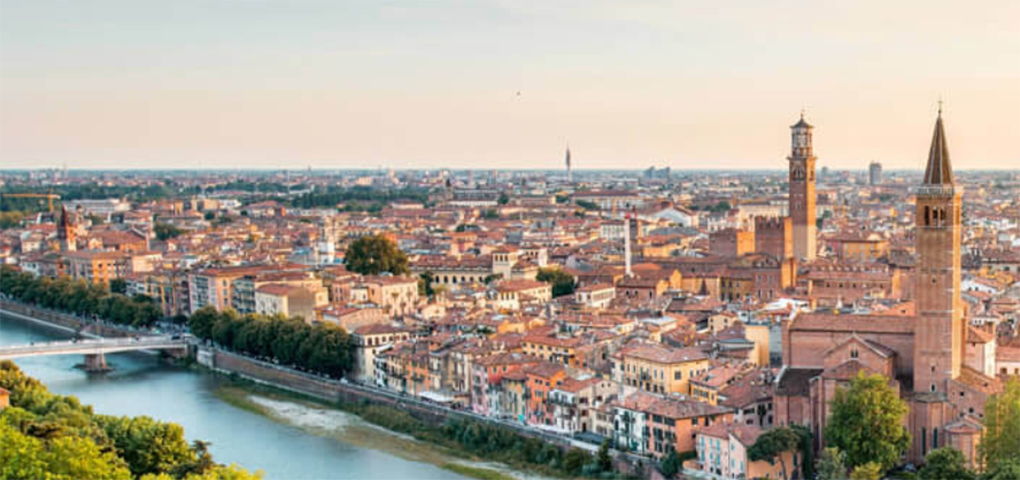
World Class Evaluations
Interbull and the Interbull Centre provide world-leading independent genetic and genomic evaluations


Our guiding principles are:
- Accurate predictions
- Independence
- Timely delivery
- Documented methods and practices publicly available
- Unbiased statistics
- Comprehensive communication
- Secure data repository
 Industry Experience
Industry Experience
 360 Million Breeding Values Per Year
360 Million Breeding Values Per Year


" Interbull services are key to enhance our genomic evaluations with international data, which is very important for Swiss breeders ."
Urs Schnyder
CEO Qualitas AG
Interbull Steering Committee Chair
2025-
Partners and Accreditations
News and Events